Vibrational Spectroscopy
Everything that living things do can be understood in terms of the jigglings and wigglings of atoms (R.P. Feynman)
Part of the Molecular Photonics group investigates the structure and dynamics ("jigglings and wigglings") of complex molecular systems by means of vibrational spectroscopy, in particular time-resolved and two-dimensional infrared spectroscopy. The investigated molecular systems include
- peptides and proteins
- soft matter
- liquid water and aqueous solutions
- catalytic transition-metal complexes
In all of these systems, the molecular motions take place on many time scales, ranging from less than a picosecond to microseconds and longer. Structural dynamics taking place on such vastly different time scales are difficult to investigate with conventional structure-resolving methods, but can be probed directly using time-resolved vibrational spectroscopy.
If you are interested in doing a BSc or MSc project, send an email to s.woutersen@uva.nl
Highlights:

Easy access to nanoparticle size and structure
Robert Schmidt used Raman Diffusion-Ordered Spectroscopy to measure nanoparticle size and structure simultaneously, in a very simple way:
Innovatiefonds Noord-Holland invests in InspecT
The UvA spin-off InspectT has obtained funding from Innovatiefonds Noord-Holland to develop and validate Optical DOSY:
InspectT ontvangt financiering van het Innovatiefonds Noord-Holland

Mystery solvent reveals its secrets
Using hexafluoro-isopropanol (HFIP) as a solvent can enhance chemical reaction rates in a spectacular manner, but how this happens is still largely unknown. Investigating the hydrogen-bond structure and dynamics of HFIP sheds new light on this mystery.
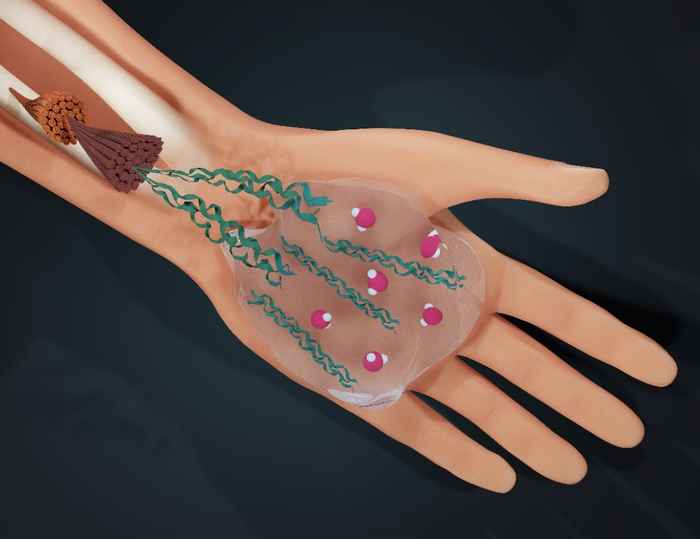
How water guides collagen self-assembly
By slightly tweaking the hydrogen bonding in water, an international team led by Giulia Giubertoni has revealed how water influences the process of collagen self-assembly.
Proof-of-Concept funding for optical DOSY
The UvA's technology-transfer office funds the development of a new spectroscopic method to determine the size of molecules and nanoparticles.
An infrared probe of molecular structure and size
Inspired by ideas from NMR, we have developed InfraRed Diffusion-Ordered SpectroscopY (IR-DOSY), which simultaneously characterizes the size and the chemical structure of a molecule, or of a mixture of molecules.
Angew. Chem. Int. Ed. 62, e202213424 (2023)
Highlight on Spectroscopy Europe
NWO Demonstrator grant for IR-DOSY
"UvA-scheikundigen bouwen een nieuw meetapparaatje voor moleculen" (article in Folia)
Adapting infrared spectroscopy for protein + amyloid mixtures
Giulia Giubertoni developed a simple experimental method to separate the infrared spectrum of an amyloid/protein mixture into its component spectra:
J. Chem. Phys. 158, 124202 (2023)
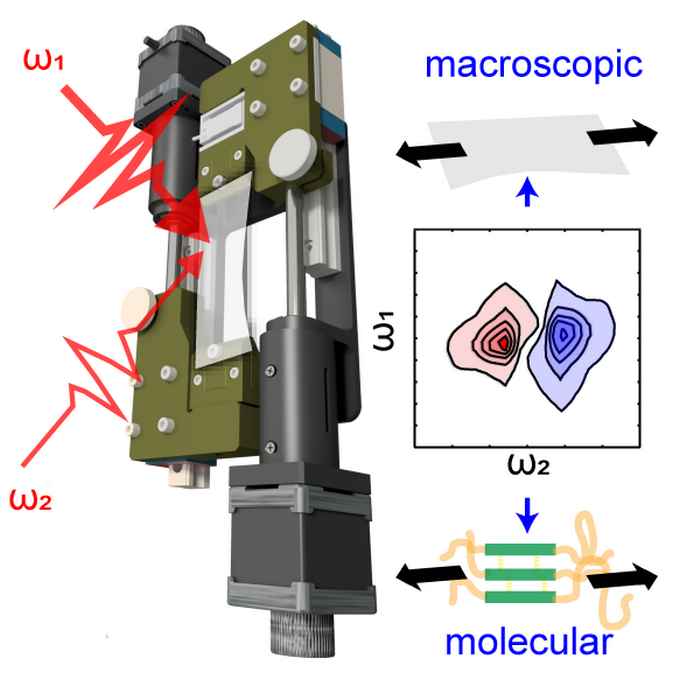
Hydrogen bonds under stress
Combining two-dimensional infrared spectroscopy with stress-strain measurements sheds new light on the mysterious elastic properties of polyurethane.
Why icicles have ribs
PhD student Menno Demmenie discovered that surface wetting is essential for the formation of ribs on icicles. He described his findings in a paper that was highlighted in Nature.

Worm chromatography
BSc student Tess Heeremans discovered that mixtures of active particles can be sorted by their activity level using a traditional chemistry method: chromatography.
Fastest rheometer in the world
Using a fluorescent molecular rotor in combination with ultrafast laser excitation, we measure liquid viscosities with nanosecond time resolution.

Ice, a self-healing material
Scratches in ice heal spontaneously over time. PhD student Menno Demmenie unraveled the physical mechanism behind this process.
Separating a mix with 2DIR spectroscopy
Peak overlap of compounds in a mixture is often a problem in vibrational spectroscopy. Postdoc Federico Caporaletti found a solution to this problem by exploiting the ultrafast relaxation of the vibrational modes. This made it possible to study the unique behavior of water in supercooled sorbitol solutions.
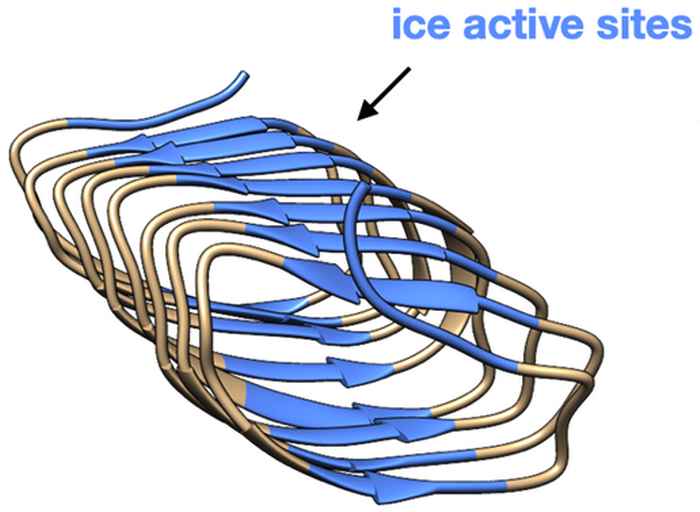
How ice-nucleating proteins influence the structure of water
A combination of surface-specific and two-dimensional vibrational spectroscopy sheds new light on how some bacteria are able to induce the freezing of water.
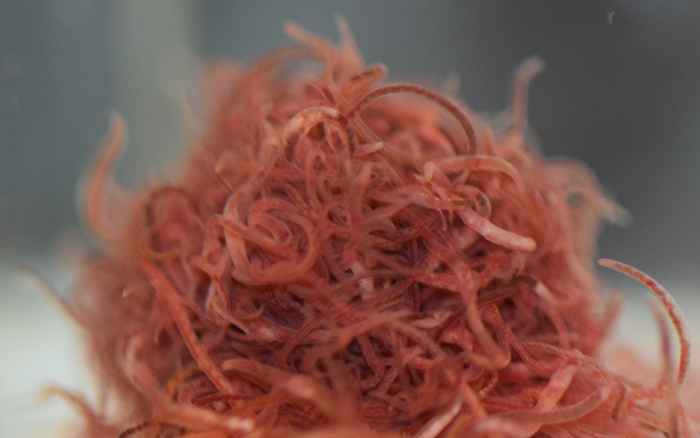

The physics of entangled worms
Active matter (ensembles of particles that consume energy in order to move or exert force) often behaves very differently from normal matter. Using a sample that we bought in the pet shop on a Friday afternoon, we found that active "polymers" show unique phase-separation and shear-thinning behaviour.
Phys. Rev. Lett. 124, 208006 (2020)
Phys. Rev. Lett. 124, 188002 (2020)
"Tipsy sludge worms simulate active polymers" (Physics World)
"Worm Viscosity" (highlight on the APS website)
"Wriggling worms help explain how certain polymers move and flow" (highlight in Chemistry World)
"UvA-onderzoek naar kluwen wormen helpt bij natuurkundige theorie" (Folia)
"The surprising viscosity of entangled worms" (highlight on the UvA website)
New light on proteins at interfaces
Many proteins are located at interfaces for their biofunctionality, which makes them problematic to study with conventional methods. Infrared sum-frequency generation provides the solution.
Fastest molecular shuttle in the world
A shuttling rotaxane synthesized by the group of Fred Brouwer uses a clever harpooning mechanism to speed up the translational motion. Matthijs Panman measured the speed using time-resolved vibrational spectroscopy.
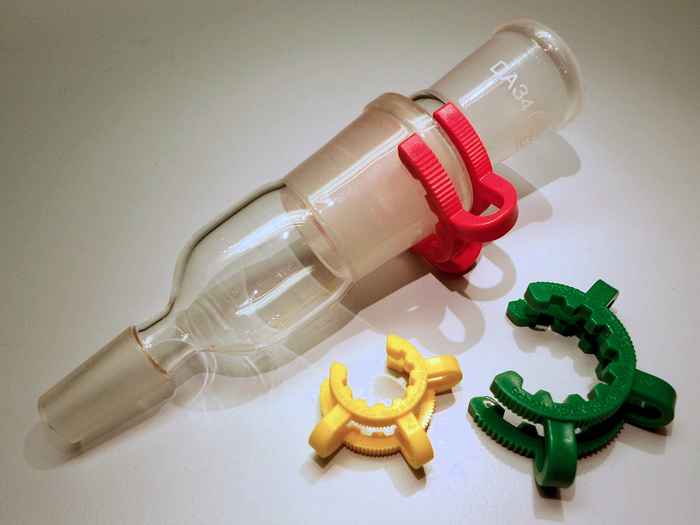
Solubility mystery solved
Why are some polyethers (such as POM, used to make Keck clips) completely insoluble in water, whereas others, such as PEG (present in nearly every cosmetic product), are perfectly miscible with water? A combination of spectroscopy experiments and computer simulations reveals the answer.
Nature Commun. 10, 2893 (2019)
"Why PEG dissolves in water but Keck clips don’t" (Chemistry World)
"Electronen delen = geen oplossing" (C2W)
"The puzzle of water solubilities of polyethers solved" (MRS Bulletin)
Van Gogh and 2D-IR spectroscopy
Using two-dimensional infrared spectroscopy, Joen Hermans has unravelled the structures of the zinc complexes involved in the degradation of zinc-white oil paint (used by Vincent van Gogh and other painters of his day). Joen's results shed new light on the humidity sensitivity of oil-paint ageing.
"The similarities between a Van Gogh painting and a golf ball" (highlight on phys.org)
Non-cooperative water dynamics in salt solutions
The isotope-dependent response of salt solutions to oscillating electrical fields shows that water molecules surrounding ions behave in a less cooperative way than they do in bulk water. These findings lead to an update of the theory for the dielectric response of salt solutions developed by Nobel Laureate Onsager and his co-worker Hubbard, and enable a reliable determination of hydration numbers, which play a key role in chemistry and biophysics.
Phys. Rev. Lett. 120, 216001 (2018)
Two kinds of deeply supercooled water
Using a combination of calorimetry, molecular dynamics simulations and infrared spectroscopy, we investigated a liquid-liquid transition in supercooled aqueous solution.
Science 359, 1127 (2018) (freely accessible PDF)
The Making Of. The story of how this cooperation started, why supercooled water does not freeze, and what happened during the experiments is told in Sander Woutersen’s inaugural lecture:
Online video of the lecture (in Dutch)
Highlights about this article:
"Scientists Watch Water Change Phase From a Liquid... to a Liquid" (Gizmodo)
"Fysici bewijzen suggestie dat er twee soorten sterk onderkoeld water zijn" (Folia)
"Evidence mounts that water exists in two liquid forms" (Chemistry World)
"The shapes of water" (phys.org)
"As formas da água: investigadora prova que a água tem dois estados líquidos" (ZAP)
Molecular bicycle pedals
A molecular machine fueled by light and operating like the crankshaft and pedals of a bicycle. With infrared pulses you can see it working in real time.
Structural snapshots of an operating molecular machine
Transient two-dimensional infrared spectroscopy makes it possible to observe the structural changes of a translational molecular machine during its operation.
A new twist to amyloid fibrils
The pathogenesis of Parkinson's disease is believed to involve the self-assembly of alpha-synuclein into amyloid fibrils. Surprisingly, when its last 32 amino acid are truncated alpha-synuclein forms much more strongly twisted fibrils.
J. Am. Chem. Soc. 139, 15392 (2017)
"Stronger twist to cytotoxic amyloid fibrils" (Science Daily)
Is "biological water" the same as tap water?
Everyone knows that plants and animals consist mostly of water. Does this water behave the same as normal tap water? To find out, we used time-resolved vibrational spectroscopy and dielectric-relaxation spectroscopy.
Nature Communications 8, 904 (2017)
"Zit er zoiets als levenswater in levende cellen?" (Volkskrant)
"Biologisch water is gewoon water" (C2W)
Highlight in Amsterdam Science magazine
"Existenz des vierten Aggregatzustandes widerlegt" (Analytik News)
Measuring the tilt angle of a protein
The absolute orientation of interfacial proteins can be determined using phase-resolved sum frequency generation spectroscopy in combination with molecular dynamics simulations and spectral calculations.
The secrets of a fungal 'raincoat'
A combination of interfacial spectroscopy with spectral calculations reveals the molecular properties of extremely water-repelling hydrophobin films.
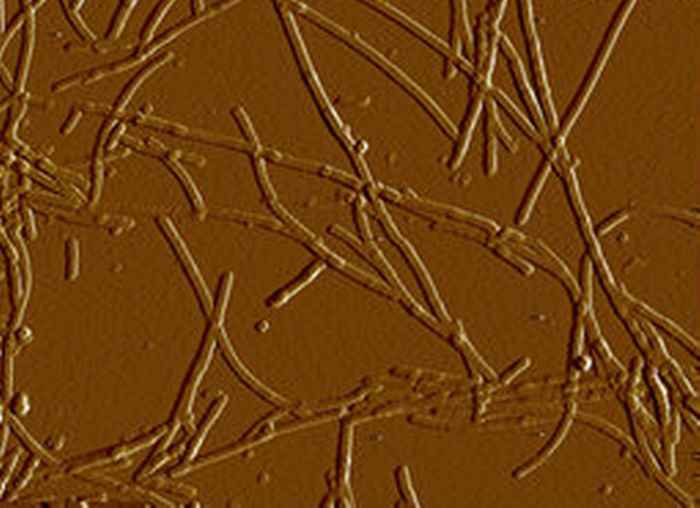
New light on Parkinson's
The protein alpha-synuclein forms fibrillar aggregates that play a key role in the pathogenesis of Parkinson's disease. The morphology of the fibrils changes dramatically depending on the amount of salt in the surrounding buffer. 2D-IR spectroscopy reveals the underlying molecular mechanism.
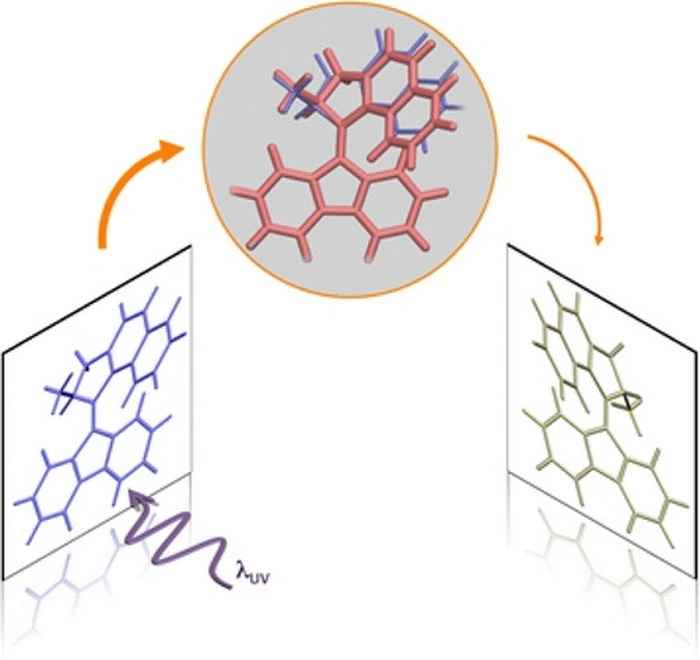
Surprising finding under the hood of molecular motors
Time-resolved experiments on the molecular motor of Nobel laureate Ben Feringa show that its operation cycle involves an electronic 'dark state' that results in a small but important stutter in the rotational motion.
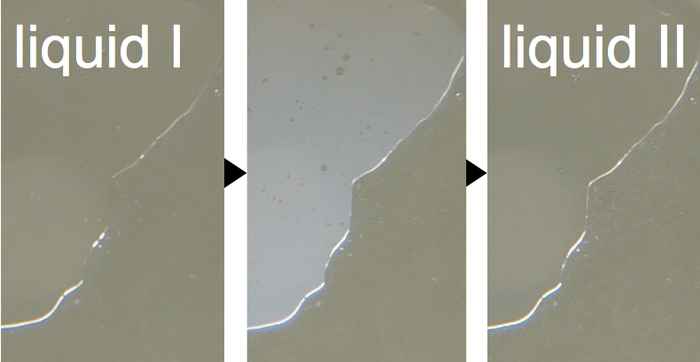
Solving water mysteries
Water containing antifreeze (glycerol) can be cooled down all the way to -100°C without freezing. At that temperature, the liquid undergoes a phase transition and changes into… another liquid?! 2DIR Spectroscopy reveals what is going on at the molecular level.

The redox behavior of a bio-inspired hydrogen-production catalyst
The Homogeneous Catalysis group of HIMS has achieved highly efficient hydrogen production using a synthetic catalyst that mimics the design of the iron-iron hydrogenase enzyme. The unique redox behavior of the catalyst was unraveled using time-resolved IR spectroscopy.

Cover story on amplified vibrational circular dichroism
Review article by Sergio Domingos highlighted on the cover:
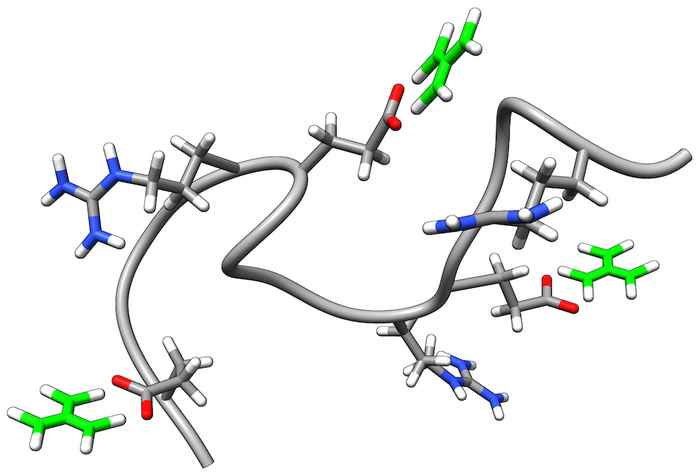
How guanidinium unfolds proteins
Guanidinium is a commonly used denaturant, but the mechanism by which it unfolds proteins is still largely unknown. We find that guanidinium disrupts the folded conformation by breaking salt bridges. 2D-IR spectroscopy shows that guanidinium binds to the carboxylate side groups involved in these salt bridges.

Switchable Amplification of Vibrational Circular Dichroism
Adding a ferrocene-based electrochemically switchable amplifier to a biomolecular system enables localized amplification of vibrational circular dichroism, making it possible to probe chirality in a site-specific manner.
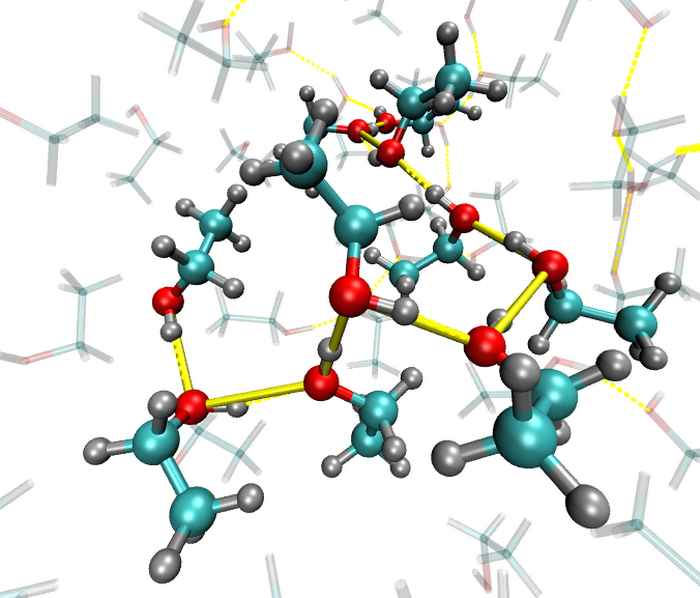
Local orientational order in liquids
Ultrafast IR spectroscopy combined with molecular dynamics simulations shows that hydrogen-bonded liquids such as ethanol and N-methylacetamide are less 'random' that you might think.
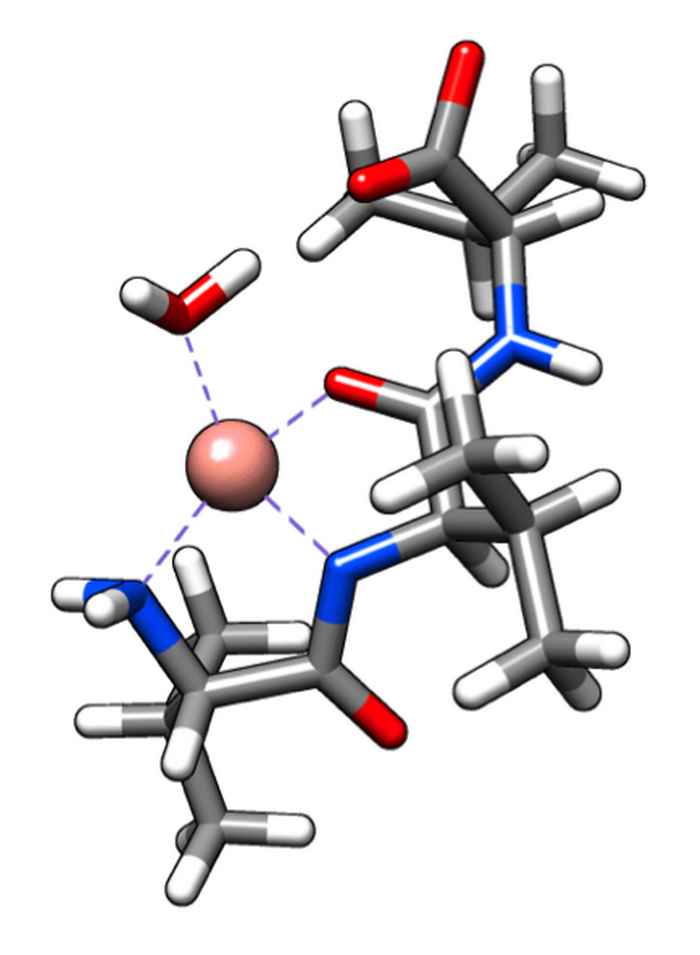
Amplifying the optical effects of chirality
The vibrational circular dichroism of amino acids and oligopeptides can be enhanced by up to 2 orders of magnitude by coupling them to a paramagnetic metal ion.
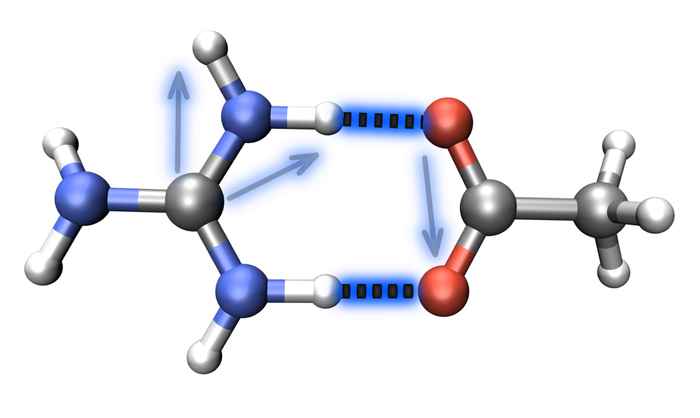
Unraveling the structure of salt bridges in solution
Salt-bridge geometries can be determined by measuring the couplings between vibrations of the salt-bridged moieties using 2D-IR spectroscopy.