Characterization of the SARS-CoV-2 Spike Receptor-Binding Domain by HILIC-MS
Collaborative work of HIMS Analytical Chemists with Pacific Northwest National Laboratory (USA)
21 April 2022
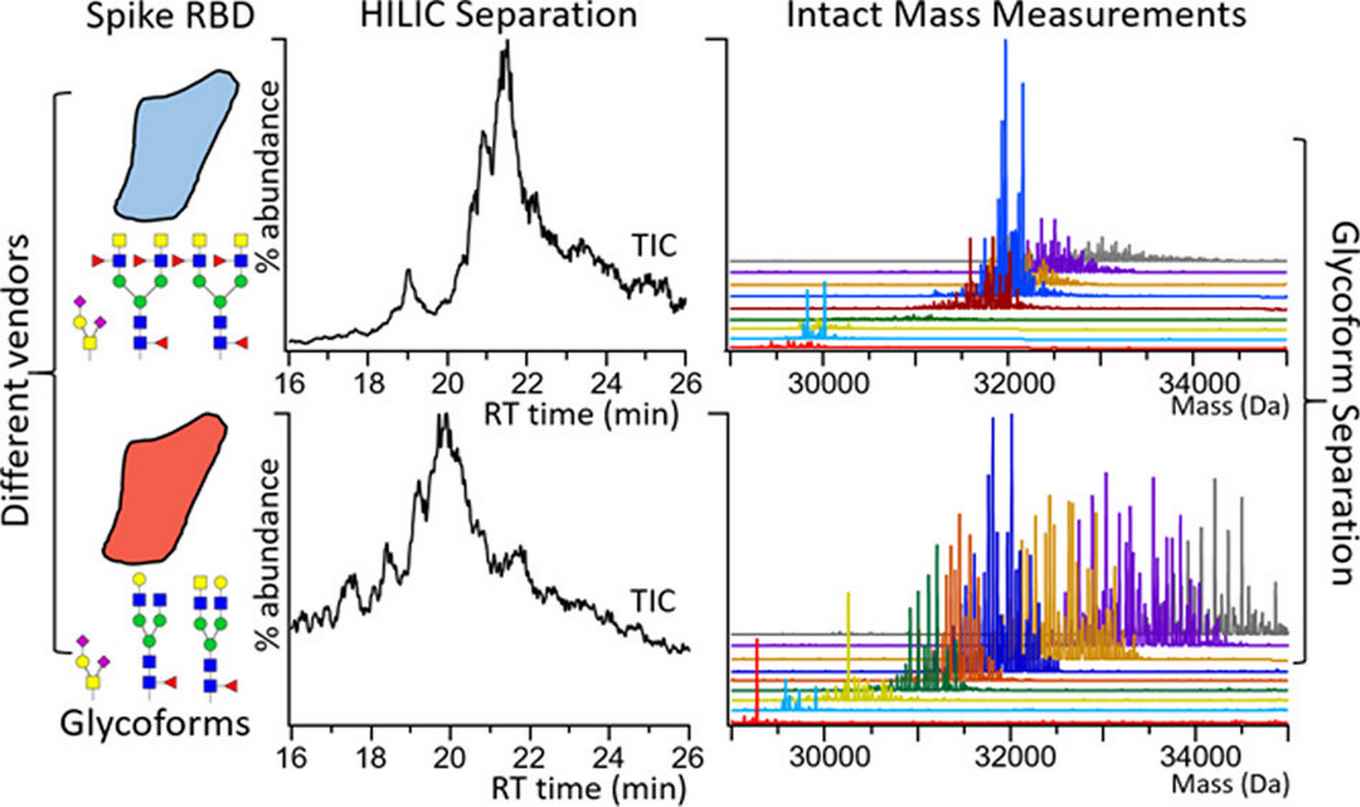
The results presented in the paper demonstrate the unique capability of the acrylamide based materials developed by Marta Passamonti during her PhD research. These allow to identify hundreds of glycoforms that were not otherwise detected by reference methods (CE and RPLC-MS). The work of Passamonti was published last year in a paper in Analytical Chemistry.
Abstract
SARS-CoV-2 cellular infection is mediated by the heavily glycosylated spike protein. Recombinant versions of the spike protein and the receptor-binding domain (RBD) are necessary for seropositivity assays and can potentially serve as vaccines against viral infection. RBD plays key roles in the spike protein’s structure and function, and thus, comprehensive characterization of recombinant RBD is critically important for biopharmaceutical applications. Liquid chromatography coupled to mass spectrometry has been widely used to characterize post-translational modifications in proteins, including glycosylation. Most studies of RBDs were performed at the proteolytic peptide (bottom-up proteomics) or released glycan level because of the technical challenges in resolving highly heterogeneous glycans at the intact protein level. Herein, we evaluated several online separation techniques:
- C2 reverse-phase liquid chromatography (RPLC);
- capillary zone electrophoresis (CZE); and
- acrylamide-based monolithic hydrophilic interaction chromatography (HILIC)
to separate intact recombinant RBDs with varying combinations of glycosylations (glycoforms) for top-down mass spectrometry (MS). Within the conditions we explored, the HILIC method was superior to RPLC and CZE at separating RBD glycoforms, which differ significantly in neutral glycan groups. In addition, our top-down analysis readily captured unexpected modifications (e.g., cysteinylation and N-terminal sequence variation) and low abundance, heavily glycosylated proteoforms that may be missed by using glycopeptide data alone.
The HILIC top-down MS platform holds great potential in resolving heterogeneous glycoproteins for facile comparison of biosimilars in quality control applications.
Publication details
Jesse W. Wilson, Aivett Bilbao, Juan Wang, Yen-Chen Liao, Dusan Velickovic, Roza Wojcik, Marta Passamonti, Rui Zhao, Andrea F. G. Gargano, Vincent R. Gerbasi, Ljiljana Pas̆a-Tolić, Scott E. Baker, and Mowei Zhou: Online Hydrophilic Interaction Chromatography (HILIC) Enhanced Top-Down Mass Spectrometry Characterization of the SARS-CoV-2 Spike Receptor-Binding Domain. Anal. Chem. 2022, 94, 15, 5909–5917. DOI: 10.1021/acs.analchem.2c00139