Novel sequencing approach enables detailed characterisation of PLGA co-polymers
15 April 2026
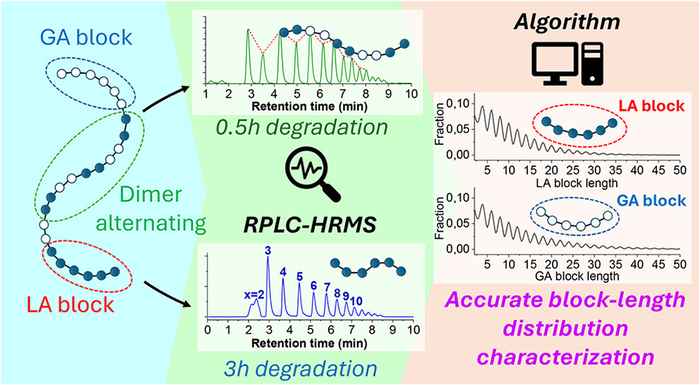
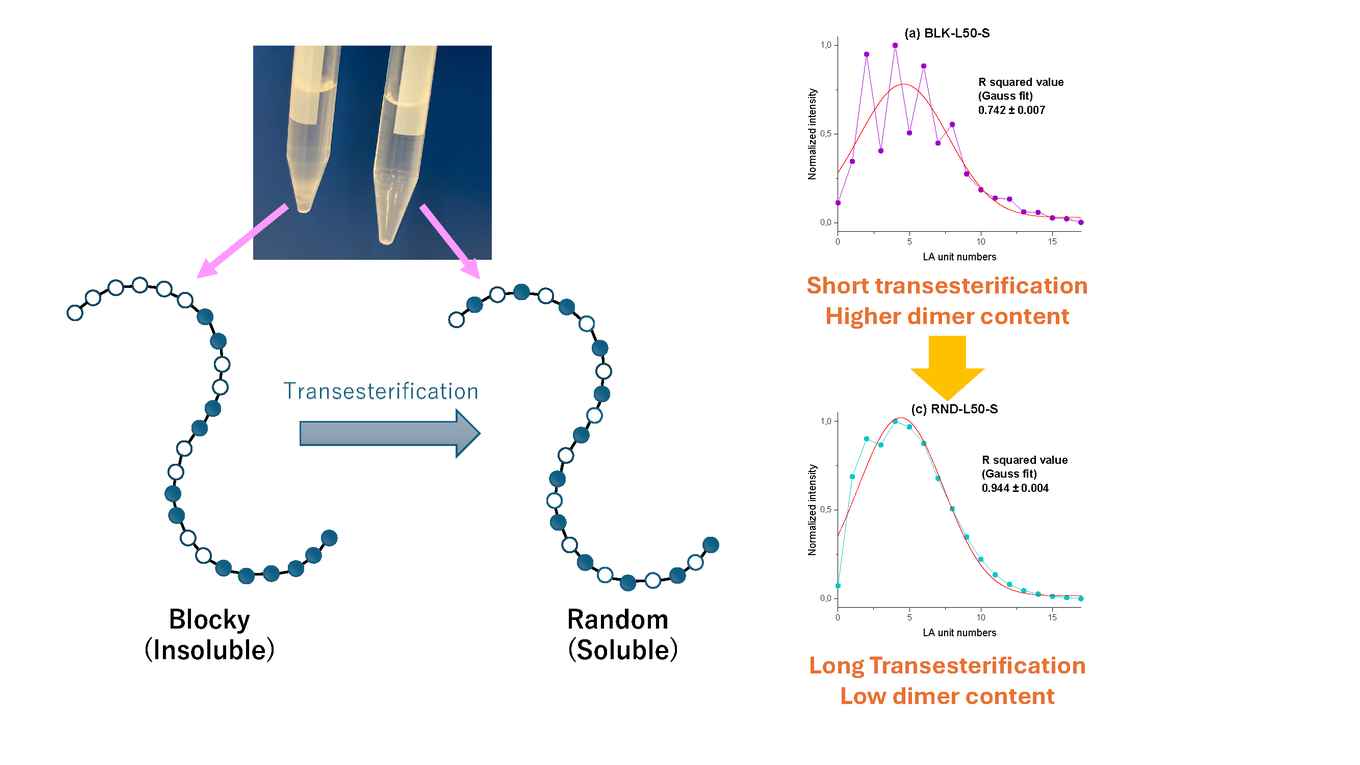
The paper in Macromolecules addresses the rather common issue that batches of poly(lactic-co-glycolic acid) (PLGA) may have the same end groups or molecular weight distribution, but differ significantly in polymer properties - which is reflected in the polymer performance in applications such as drug release. This is because current state-of-the-art methods for polymer analysis provide only limited information about the chemical composition and molecular weight distribution of the polymers.
The sequencing approach presented in the paper significantly improves this. The co-polymers are hydrolysed, releasing oligomers (e.g., up to 10 units) that can be analysed by RPLC-HRMS. Since polyGA displays faster hydrolysis, the resulting oligomer intensity varies between blocky and random PLGA. The paper presents a method to reconstruct the block length of the polymer from dedicated mass spectrometry (SWAMP-MS) data, which was developed in cooperation with colleagues Tijmen Bos and Bob Pirok at the Chemometrics and Advanced Separations Team (CAST, part of the HIMS Analytical Chemistry group).
Proof of principle
The research forms part of the PhD research of Masashi Serizawa, supervised by Gargano and sponsored by Mitsubishi Chemicals. The PLGA project was supported through a 50,000 euro TAFU-XL grant from COAST (the Community of Innovation for Comprehensive Analytical Science and Technology), which entailed cooperation with the company Corbion.
Now that proof-of-principle has been established, Gargano hopes to further develop the method in collaboration with industry and launch subsequent public-private-partnership (PPP) grant applications.
Paper details
Masashi Serizawa, Dion Snijders, Stefan van Berkel, Pieter van Delft, Tijmen S. Bos, Bob W. J. Pirok, Ron A.H. Peters, Peter J. Schoenmakers, and Andrea F.G. Gargano: Quantifying Poly(lactide-co-glycolide) Residual Dimer-Alternating Structure and Block-Length Distribution Using Controlled Chemical Degradation and Reverse-Phase Liquid Chromatography–Mass Spectrometry Analysis Macromolecules, Article ASAP, DOI: 10.1021/acs.macromol.5c03061